fastqq is intended for creating quantile-quantile plots.
We provide faster alternatives to qqman::qq,
stats::qqplot and stats::qqnorm. We also
provide the function fastqq::drop_dense such that the user
can extract the data to plot with ggplot.
For 100 million samples, we achieve 80X speedup
compared to qqman::qq. This takes the running time
from ~13.5 minutes down to less than 10 seconds.
This package was originally intended to speedup the creation of QQ plots for genome wide association studies (GWAS). Then I decided to make it a general tool fro QQ style plots. For QQ plots, the user often plots tens to hundreds of millions of points. Creating scatter plots with so many points is usually not efficient since the graphics devices store all the data, such that the visualization can be rescaled or plotted in a vector graphics format (where again all the data is stored).
A better and faster approach in these cases is to note that many of the points are so close to each other that there is no value in including them in the plot. QQ style plots are usually a monotonically increasing sequence of points, so we can easily employ fast filtering to remove redundant points, that would otherwise not be visible in the final plot anyways.
See examples below.
Note that this package is inspired by the
qqman package, which has now been archived. The
interface to the qq function should be very similar, and
fastqq::qq is a drop-in replacement for
qqman::qq. I created this package since it could take more
than 10 minutes to render a single plot with qqman::qq.
This is also to save on memory and other resources, in particular
time.
You can install the released version of fastqq from CRAN with:
install.packages("fastqq")And the development version from GitHub with:
# install.packages("devtools")
devtools::install_github("gumeo/fastqq")The following is an example from very simple simulated data:
suppressPackageStartupMessages(library(fastqq))
set.seed(42)
p_simulated <- runif(1e5)
# Classic way to do this with qqman
qqman::qq(p_simulated)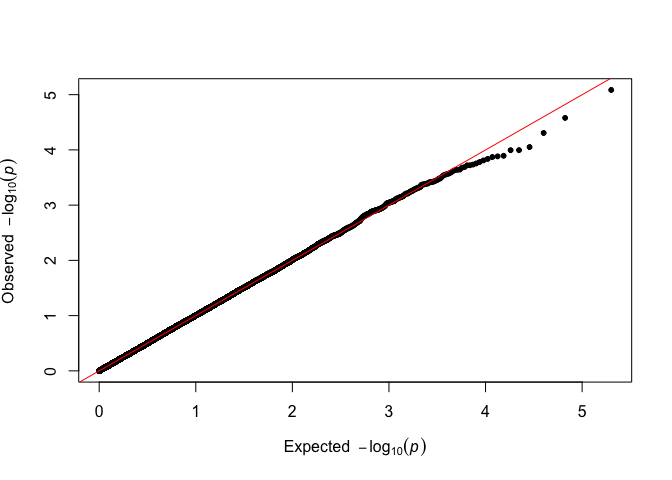
# Alternative
fastqq::qq(p_simulated)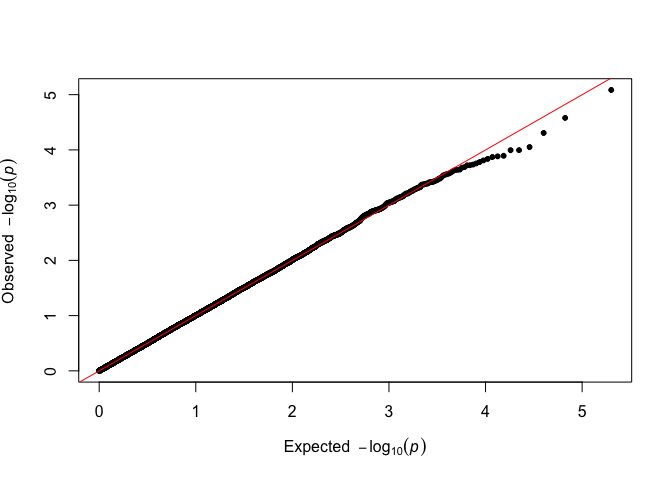
There is no visible difference, and the analysis can proceed as usual.
If some p-values are exactly zero (e.g. from software that reports
underflowed results), use the zero_action parameter to
substitute a small finite value:
pvec_with_zeros <- c(runif(1e4), 0, 0)
qq(pvec_with_zeros, zero_action = 1e-300)
#> Warning in qq(pvec_with_zeros, zero_action = 1e-300): 2 p-value(s) equal to
#> zero replaced with 1e-300.
We can compare the timings of creating the plots, with
qqman.
set.seed(555)
N_test <- c(1e3,1e4,1e5,1e6)
time_method <- function(pkg_name, method){
suppressPackageStartupMessages(library(pkg_name,
character.only=TRUE, quietly = TRUE))
for(N in N_test){
p_vec <- runif(n = N)
print(paste0("Timing ", pkg_name, "::", method," with ",
N, " points"))
tictoc::tic()
pdf(file = NULL) # Prevent the plots from appearing
do.call(method, list(pvector=p_vec))
dev.off()
tictoc::toc()
}
}
N_test <- c(1e3,1e4,1e5,1e6,1e8)
time_method('fastqq','qq')
#> [1] "Timing fastqq::qq with 1000 points"
#> 0.028 sec elapsed
#> [1] "Timing fastqq::qq with 10000 points"
#> 0.02 sec elapsed
#> [1] "Timing fastqq::qq with 1e+05 points"
#> 0.049 sec elapsed
#> [1] "Timing fastqq::qq with 1e+06 points"
#> 0.414 sec elapsed
#> [1] "Timing fastqq::qq with 1e+08 points"
#> 48.603 sec elapsed
N_test <- c(1e3,1e4,1e5,1e6)
time_method('qqman','qq')
#> [1] "Timing qqman::qq with 1000 points"
#> 0.002 sec elapsed
#> [1] "Timing qqman::qq with 10000 points"
#> 0.02 sec elapsed
#> [1] "Timing qqman::qq with 1e+05 points"
#> 0.189 sec elapsed
#> [1] "Timing qqman::qq with 1e+06 points"
#> 1.989 sec elapsedSo we can expect around 25X speedup for a million points.
For 100 million points (order of magnitude for modern GWAS),
fastqq::qq takes 10 seconds on the same hardware as for the
timings above, qqman::qq takes more than 13.78 minutes for
100 million points (80X speedup) and if one saves to a vector
graphic output, all the data is stored, and the file size scales with
the amount of points.
qqlog exampleqqlog accepts pre-computed −log₁₀(p-values). This is
useful when you have already transformed your p-values, or need full
precision before plotting (e.g. values below
.Machine$double.xmin computed in extended precision):
set.seed(42)
suppressPackageStartupMessages(library(fastqq))
pvec <- runif(1e5)
fastqq::qqlog(-log10(pvec))
qqchisq1 exampleqqchisq1 accepts χ² test statistics and converts them to
−log₁₀(p) assuming 1 degree of freedom. The conversion uses log-space
arithmetic for numerical stability even for very large statistics:
set.seed(42)
suppressPackageStartupMessages(library(fastqq))
chisq_vals <- rchisq(1e5, df = 1)
fastqq::qqchisq1(chisq_vals)
A χ² value of ~2303 (df=1) corresponds to p ≈ 10⁻⁵⁰⁰, which is far
below R’s double-precision floor of .Machine$double.xmin ≈
2.2×10⁻³⁰⁸. Naive computation underflows to zero, making the point
invisible to any method that works on the raw p-value scale:
suppressPackageStartupMessages(library(fastqq))
extreme_chisq <- 2303 # p ≈ 10^-500
# Naive approach: p-value underflows to 0, point is lost
cat("Naive p-value: ", pchisq(extreme_chisq, 1, lower.tail = FALSE), "\n")
#> Naive p-value: 0
# Log-space gives the correct -log10(p) even for such extreme values
cat("-log10(p) via log-space:",
-pchisq(extreme_chisq, 1, lower.tail = FALSE, log.p = TRUE) / log(10), "\n")
#> -log10(p) via log-space: 501.8695
# qqchisq1 uses log-space internally, so the extreme point appears correctly
set.seed(42)
chisq_vals <- c(rchisq(1e4, df = 1), extreme_chisq)
fastqq::qqchisq1(chisq_vals)
The extreme point appears at the top of the plot (~500 on the y-axis) without any special handling required from the user.
qqnorm exampleWe can use qqnorm just like from
stats::qqnorm. The only difference is in the output, we
return sorted output, and exclude NAs.
set.seed(42)
suppressPackageStartupMessages(library(fastqq))
fastqq::qqnorm(rnorm(1e6))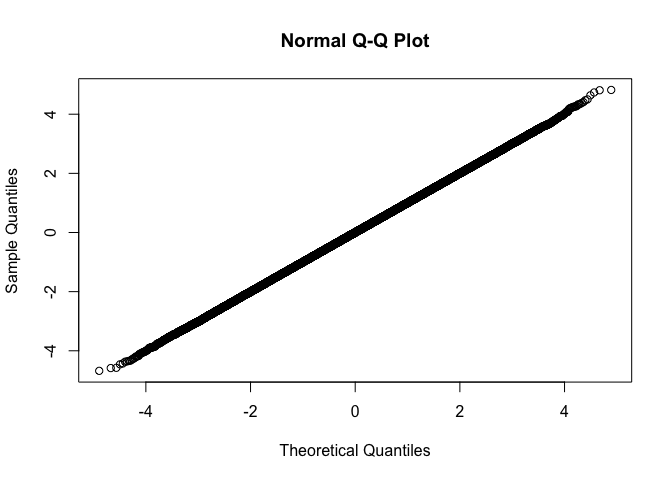
qqplot exampleset.seed(42)
suppressPackageStartupMessages(library(fastqq))
fastqq::qqplot(rnorm(1e6),rnorm(1e6))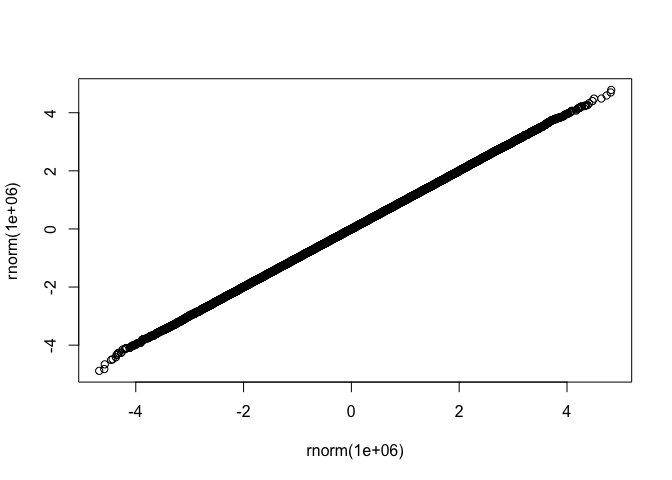
drop_dense
and plot with ggplot exampleset.seed(42)
suppressPackageStartupMessages(library(fastqq))
suppressPackageStartupMessages(library(ggplot2))
x <- rnorm(1e6)
y <- rnorm(1e6)
df <- fastqq::drop_dense(x, y)
ggplot(df, aes(x=x,y=y)) + geom_point()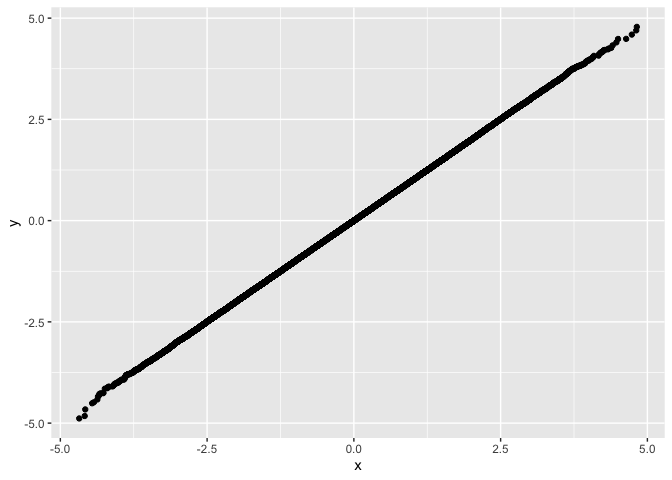
After I created this, I have found several sources, that aim at something similar, usually also a manhattan plot (I am probably also missing other packages):
fastman
package. Uses scattermore, so the plotting is very fast.
This package is not currently (31/07/2021) on CRAN.ramwas
package. Has not been maintained in 2 years and is on bioconductor. This
package is aimed for Fast Methylome-Wide Association Study
Pipeline for Enrichment Platforms and the
ramwas::qqPlotFast function is just a minor part of the
package. ItHere are also some projects on CRAN, where the plotting is similar to
qqman::qq, and is not improved for speed.
gwaRs
package. Focuses on using ggplot.manhplot
package. This is a pretty ambitious project, with a good publication. Again,
the main focus is on the manhattan plot.qqman
package. Probably one of the main inspiration for most of the other
packages.CMplot
package. Great circular manhattan plot.